I reflect on my past experiences with event-based simulation, connecting it to Queuing Theory and the statistical mathematics used in telephone exchanges and modern technological applications, particularly gaming platforms like Zwift. I draw parallels between call arrival patterns and infection models, particularly regarding COVID-19. I acknowledge the influence of historical research, including work by Prof. Neil Ferguson, on understanding infection dynamics and the necessity for public health measures like social distancing. I emphasize the importance of mathematical modeling in managing infectious diseases effectively.
Tag Archives: SARS-CoV-2
Official UK Coronavirus reporting ends
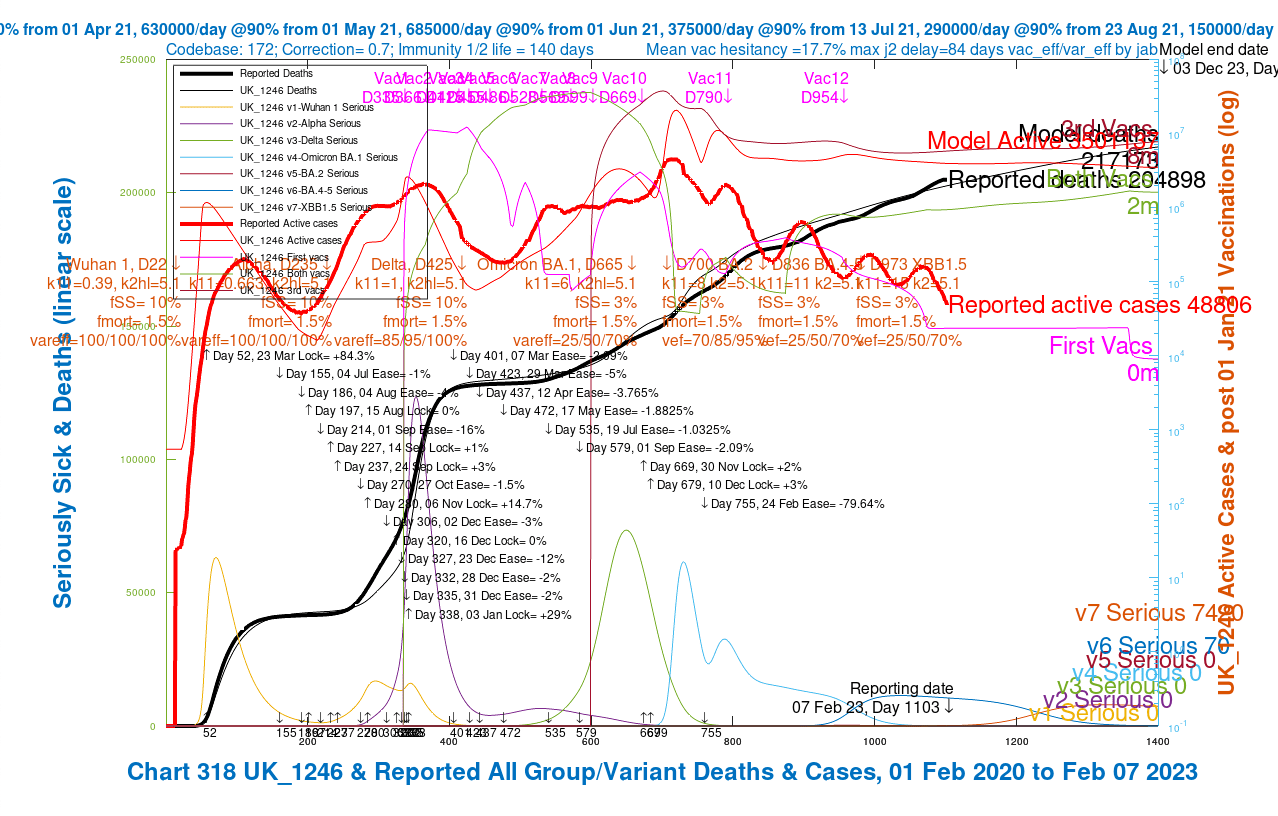
Introduction I have continued to run my own Coronavirus model, my recent prior report on February 9th this year having immediately followed the outbreak of the “Kraken” variant, Omicron XBB.1.5, in the UK. This brief post provides my own latest charts from my model, in the light of cessation of the UK Government’s own CovidContinue reading “Official UK Coronavirus reporting ends”
The Kraken Wakes* – Coronavirus
I have continued to improve and run my Coronavirus model daily. The code for the model is now at version 172, and I have added a dataset to represent the variant XBB.1.5, known informally as “Kraken”. The fit of the model to reported deaths is good. I explore in the article the behaviour of Modelled vs. Reported Active Cases.
Omicron continues to surprise – Coronavirus
Introduction In my most recent articles on the Omicron variants of Covid-19, I have highlighted the discrepancy between plummeting Active Cases and increasing cumulative Deaths. My model draws attention to this, by comparing the previous alignment of modelled and reported data, prior to February 24th, with the recent divergence. This post continues to summarise theContinue reading “Omicron continues to surprise – Coronavirus”
Declining Omicron cases but increasing deaths – Coronavirus
Something odd seems to be happening with the relationship between reported UK Covid-19 cases and deaths in the last few weeks, when deaths have been increasing somewhat more quickly than recently, but cases have declined steeply. My model charts seem to expose this oddity.
They think it’s all over – but it isn’t – Coronavirus
In this article, I present a few variations of the current model to illustrate some outcomes depending on different assumptions about the current dominant variant, Omicron BA.2; they are fairly consistent in their forecasts over the 1400 day period of the model, from the outset in February 2020 to December 2023, and show that as NPIs are removed, vaccination is what keeps us safe.
Omicron BA.2, vaccination for 5-11 year-olds, and the honeymoon period – Coronavirus
I haven’t needed to make significant updates to my Coronavirus model for a while, because it has been working well.
The original Omicron variant morphed into the new BA.2 variant, and although it seems no more dangerous than its predecessor, it is thought to be between 33%-50% more transmissible. I have assumed the lower value of 33% more transmissive for this post.
I have added Omicron BA.2 as fifth variant v5 to my model, with 8 times the transmissibility of Delta, compared with the original Omicron variant v4 in the model, at 6 times the Delta transmission rate.
The Coronavirus pandemic situation in Europe
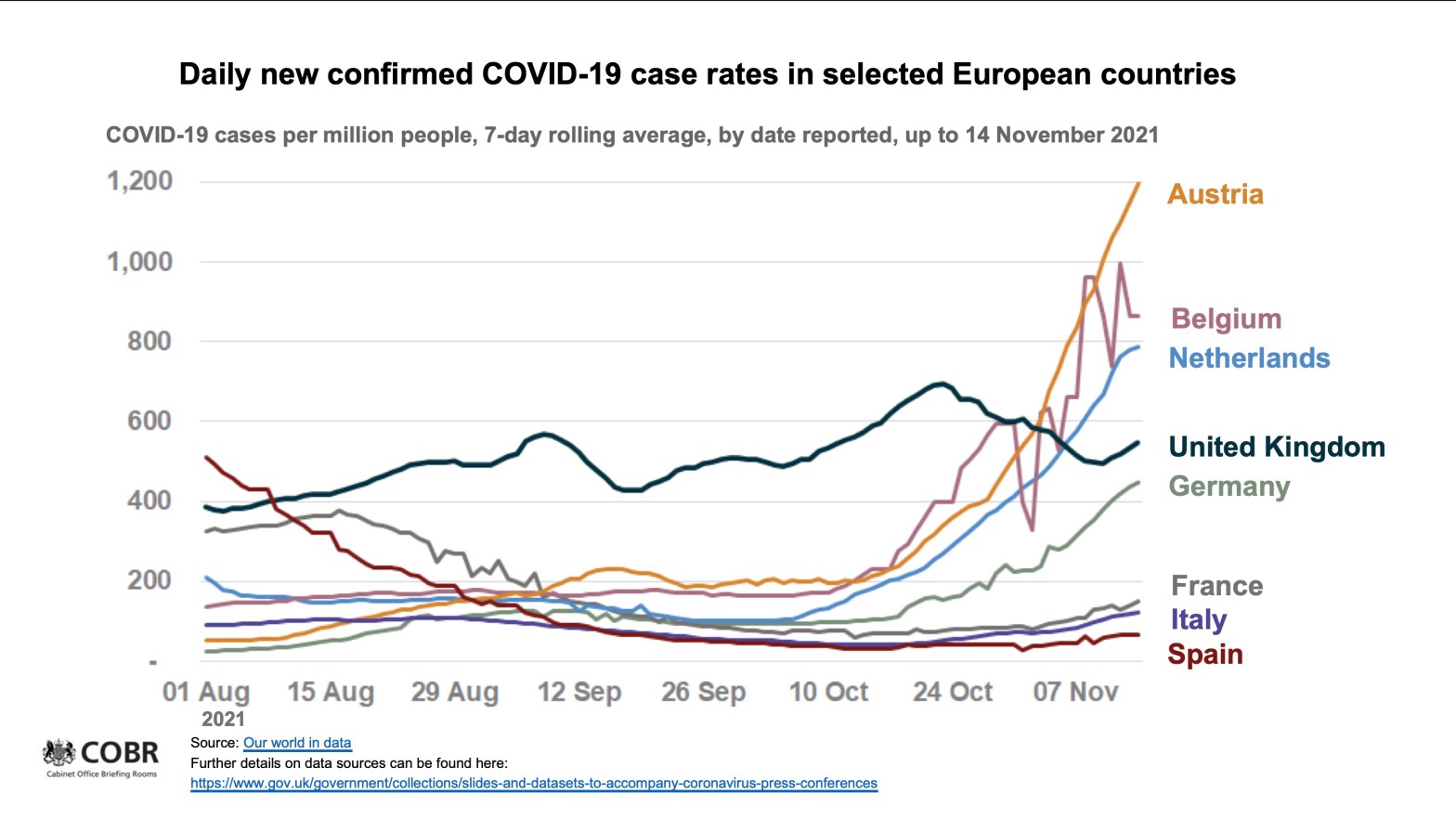
The pandemic situation in continental Europe has been worsening rapidly, and I felt that it I should update some country comparisons in a dedicated post.It confirms that the sourcing of data for a Coronavirus model of any given country is a very specific task nowadays, given the considerable differences in the underlying demographics, cultures, Government actions (NPIs) and public responses in the various countries.
This blog post isn’t looking at the modelling per se, but concentrates on the very different outcomes we are seeing across Europe, and looking at some of the reasons why.
Coronavirus pandemic modelling review
Vaccination has somewhat stabilised the SARS-Cov-2 pandemic in the UK. I summarise the capabilities that I have found necessary and useful in modelling the behaviour of the pandemic; successive variants, different population age-groups, the effect of Government NPIs, and vaccinations.
Protected: Notes for Twitter group – Coronavirus and Gompertz
There is no excerpt because this is a protected post.